The challenge of biotechnology data fragmentation
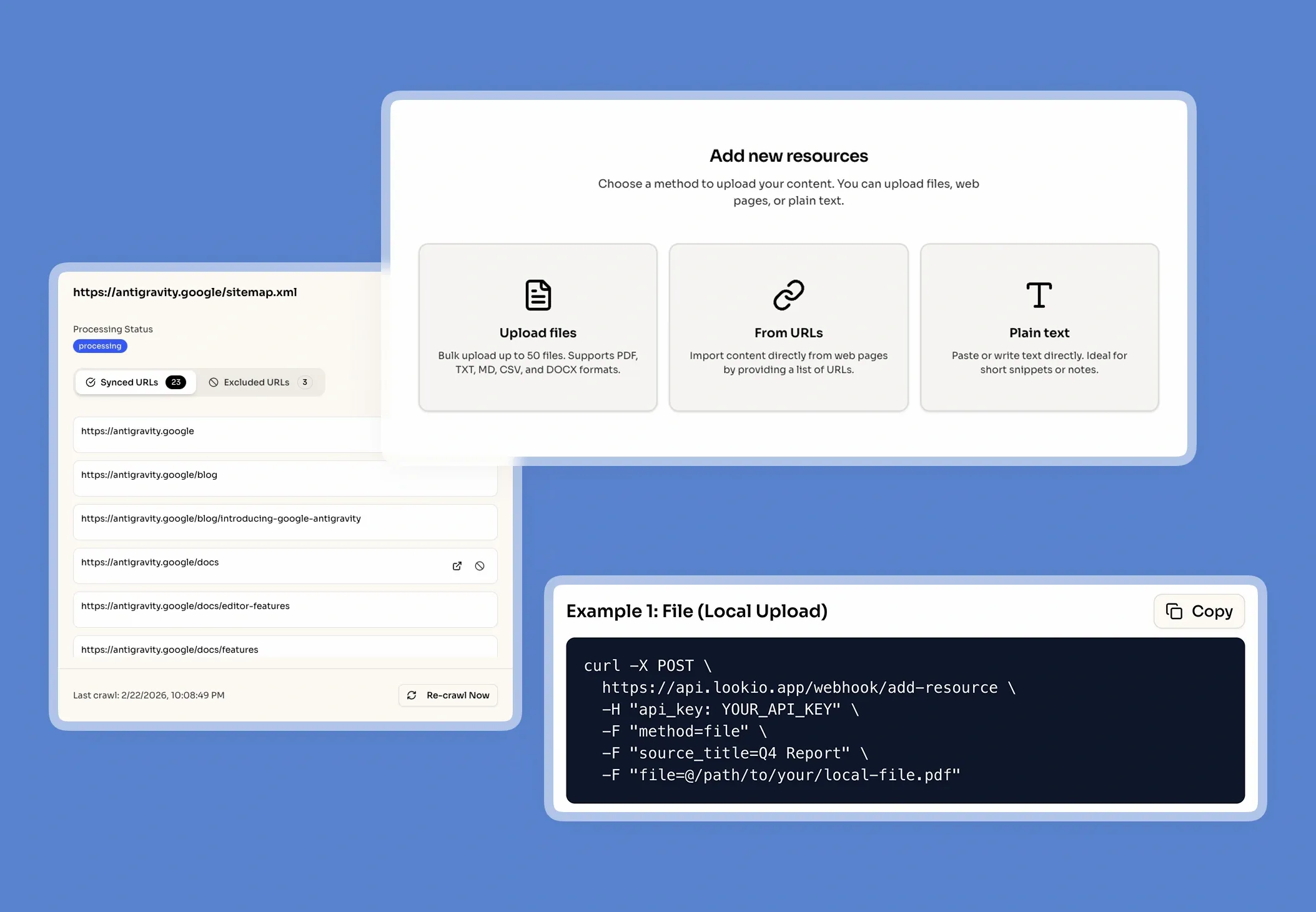
When a research scientist at a biotechnology firm needs to cross-reference a specific protein folding hypothesis against three years of internal lab notes, they often hit a digital brick wall. The data they need isn't on the public internet; it is locked inside complex PDFs, proprietary Docx files, and unstructured research logs. This fragmentation forces highly paid experts to spend hours acting as manual library filers rather than innovating.
The daily cost of the manual search
In the biotech sector, the cost of information gaps isn't just lost time—it is a risk to SLA compliance and regulatory filings. When a technical writer drafts an FDA submission and cannot instantly find the supporting trial data, the entire pipeline slows down. Every minute spent digging through a shared drive is a minute stolen from drug discovery or clinical validation. This constant interruption of subject matter experts destroys deep-work cycles and leads to inconsistent answers across the organization.
Why the tools they have tried fall short
Most biotech firms have already experimented with standard digital solutions, only to find they cannot handle the precision required for life sciences:
- Internal wikis and keyword search: These systems rely on exact word matching. If you search for a synonym or a related molecular pathway, you get zero results. They simply don't scale as documentation grows into the millions of words.
- Generic LLMs (ChatGPT): Large language models are notoriously prone to hallucination. In a field like biotechnology, an invented statistic or a misquoted chemical concentration is a liability. Furthermore, generic tools lack the security needed for sensitive IP.
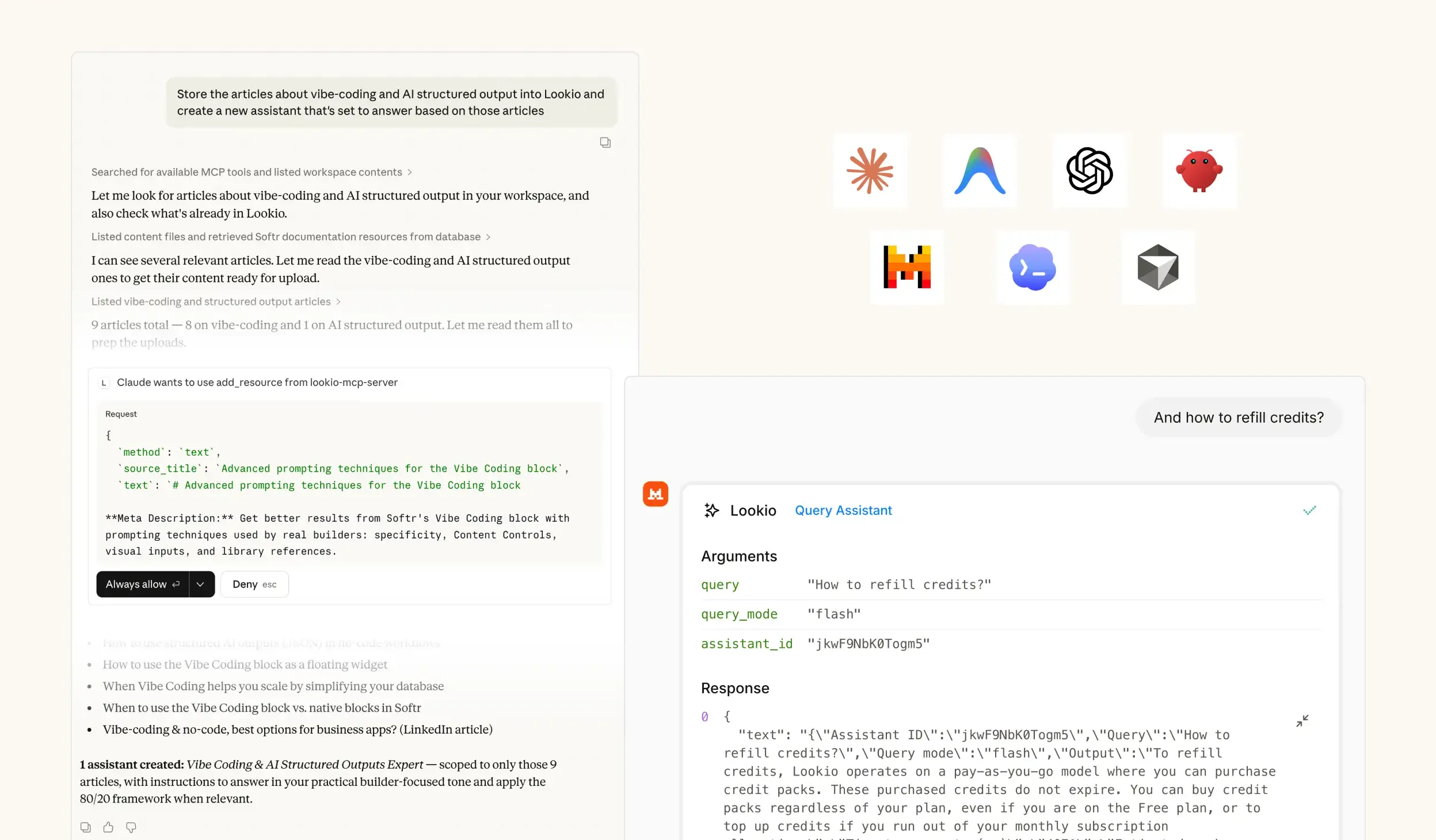
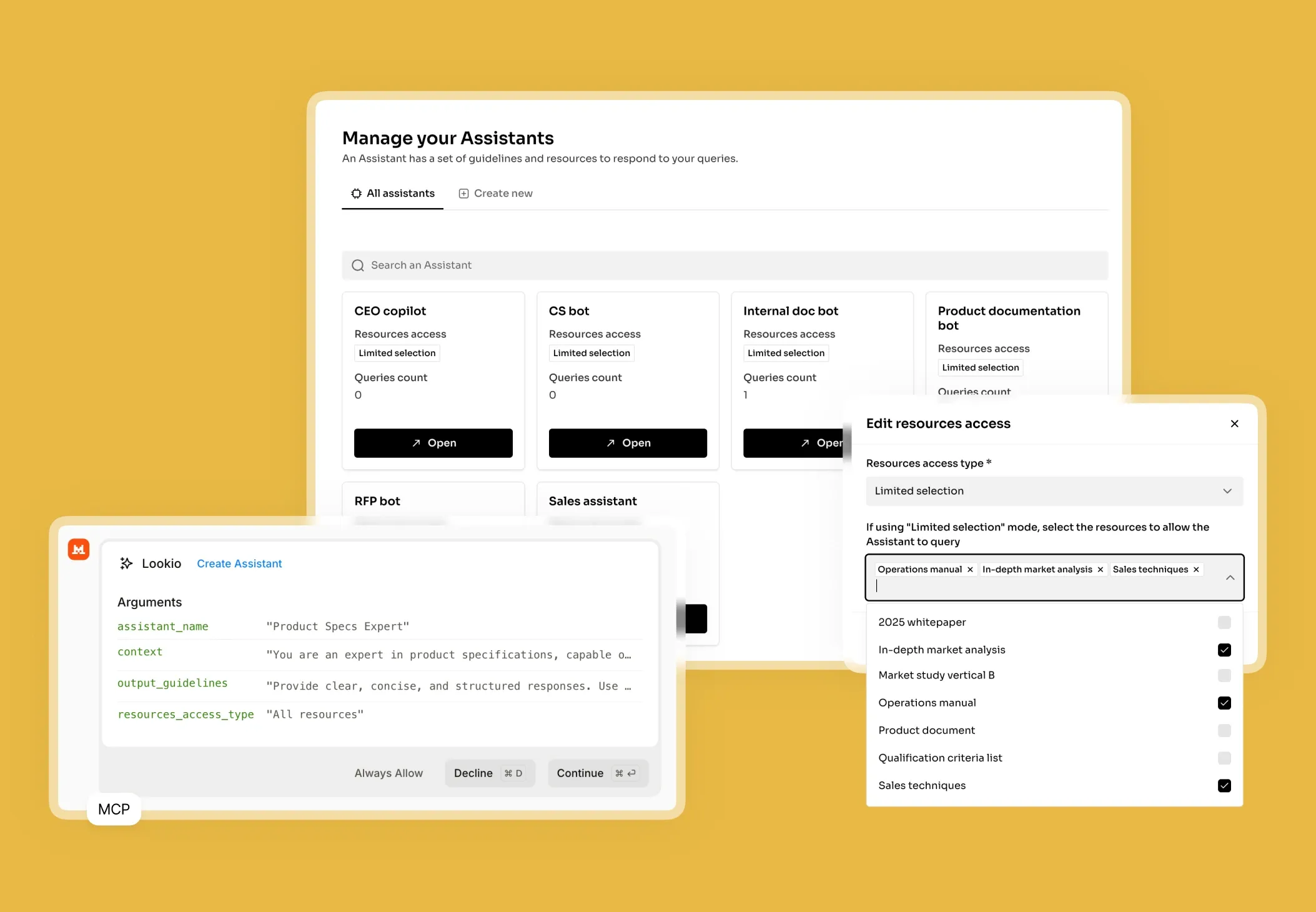
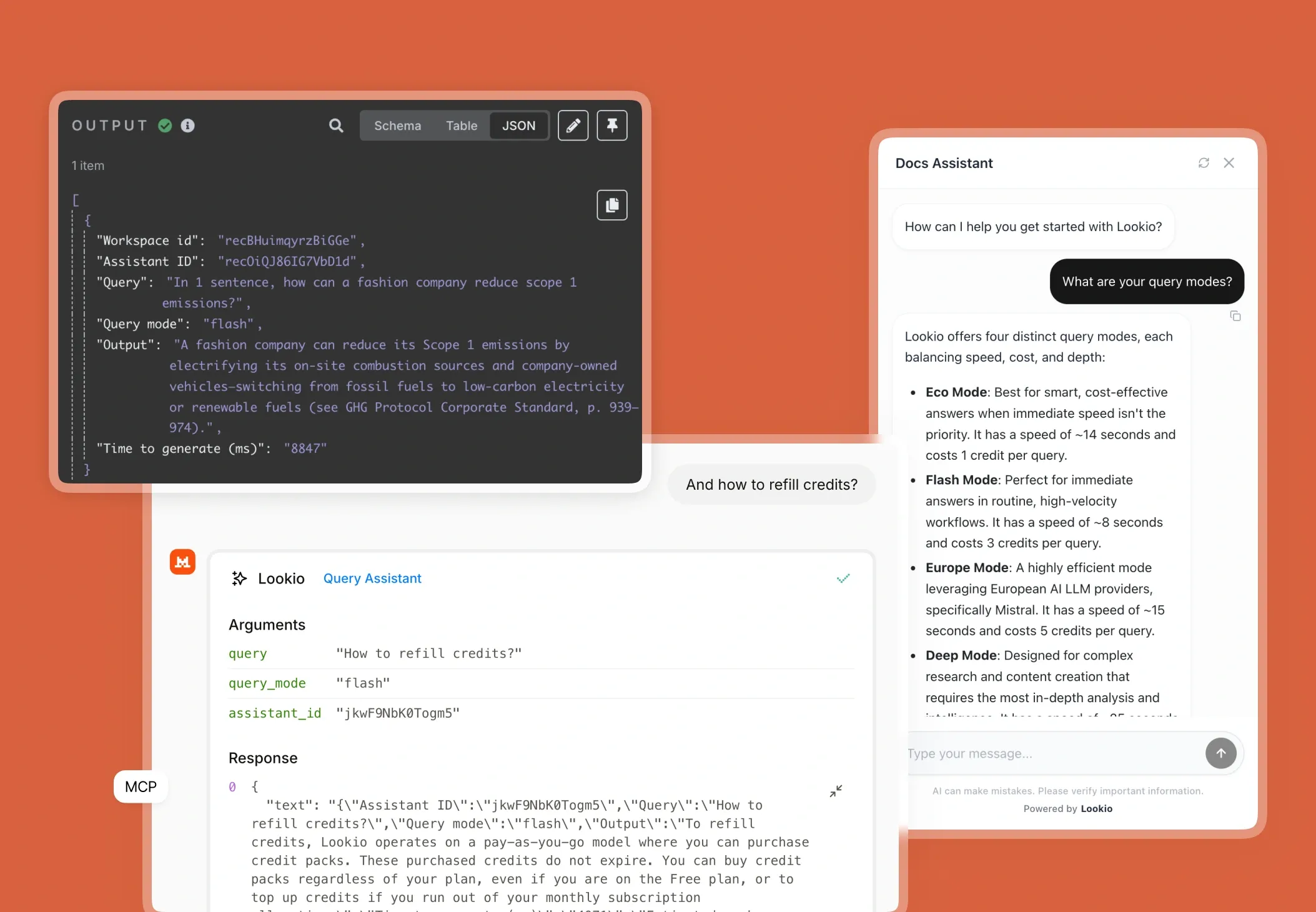
- NotebookLM and Custom GPTs: While impressive for individual research, these are not viable for business workflows because they lack a programmatic interface. You cannot connect them to your existing LIMS (Laboratory Information Management System) or automate them via tools like n8n.
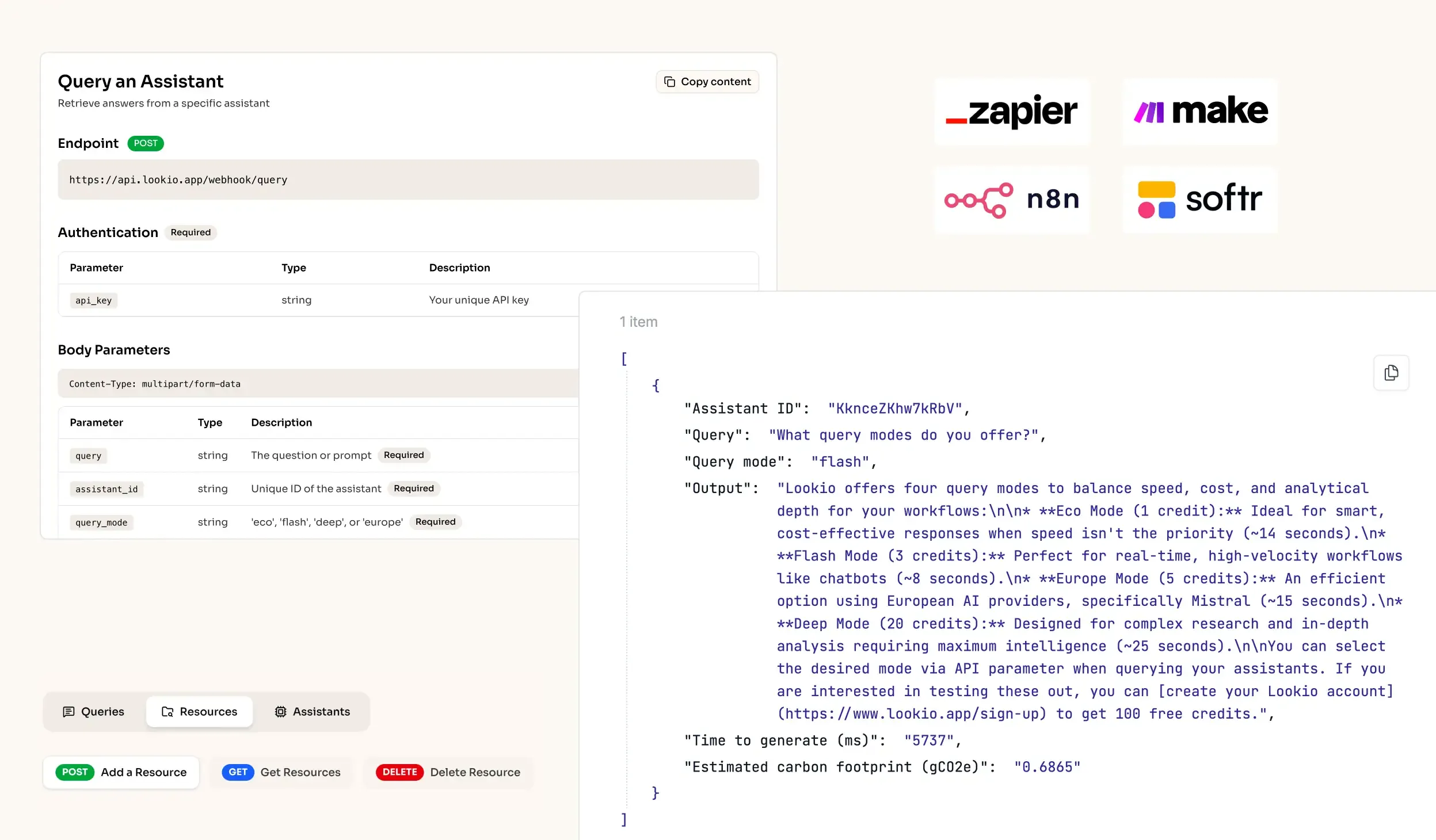
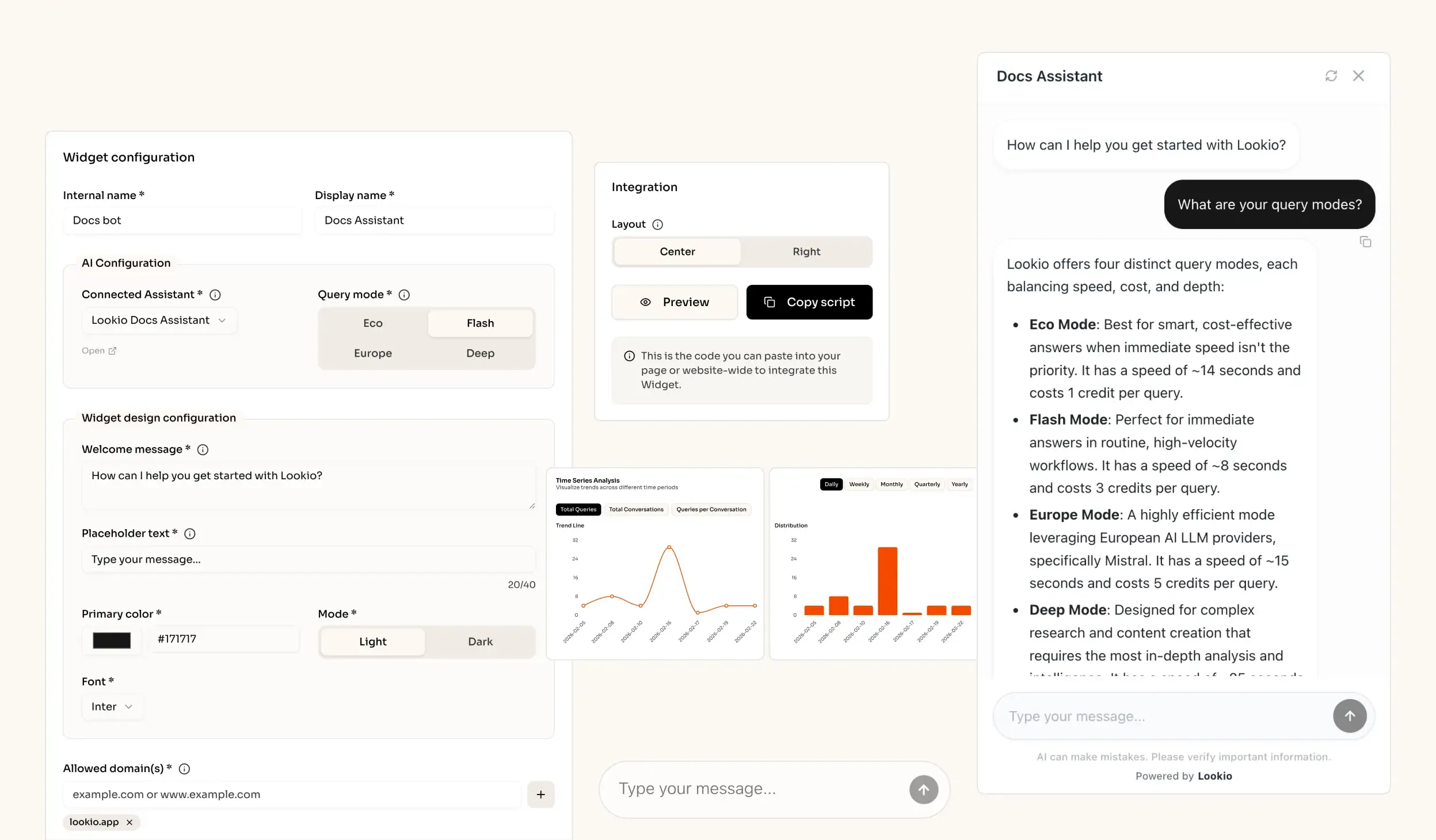
What’s missing is a way to turn static research documents into a live, queryable brain that can be accessed by both humans and automated agents alike.